Projects
Software tools and resources developed by our lab for computational biology, gene expression analysis, and metabolic network modeling.
Featured
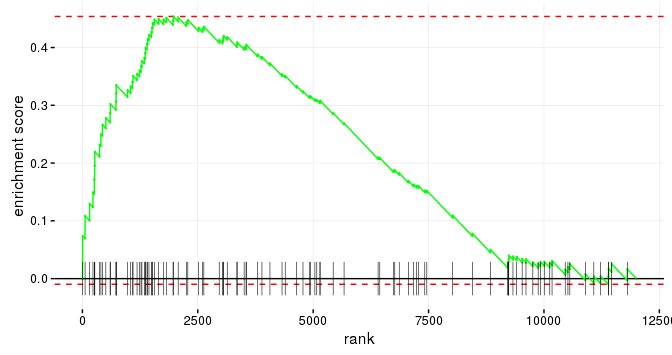
Fast gene set enrichment analysis. Implements algorithms that allow more permutations and finer-grained p-values, enabling accurate standard approaches to multiple hypothesis correction.
alserglab/fgsea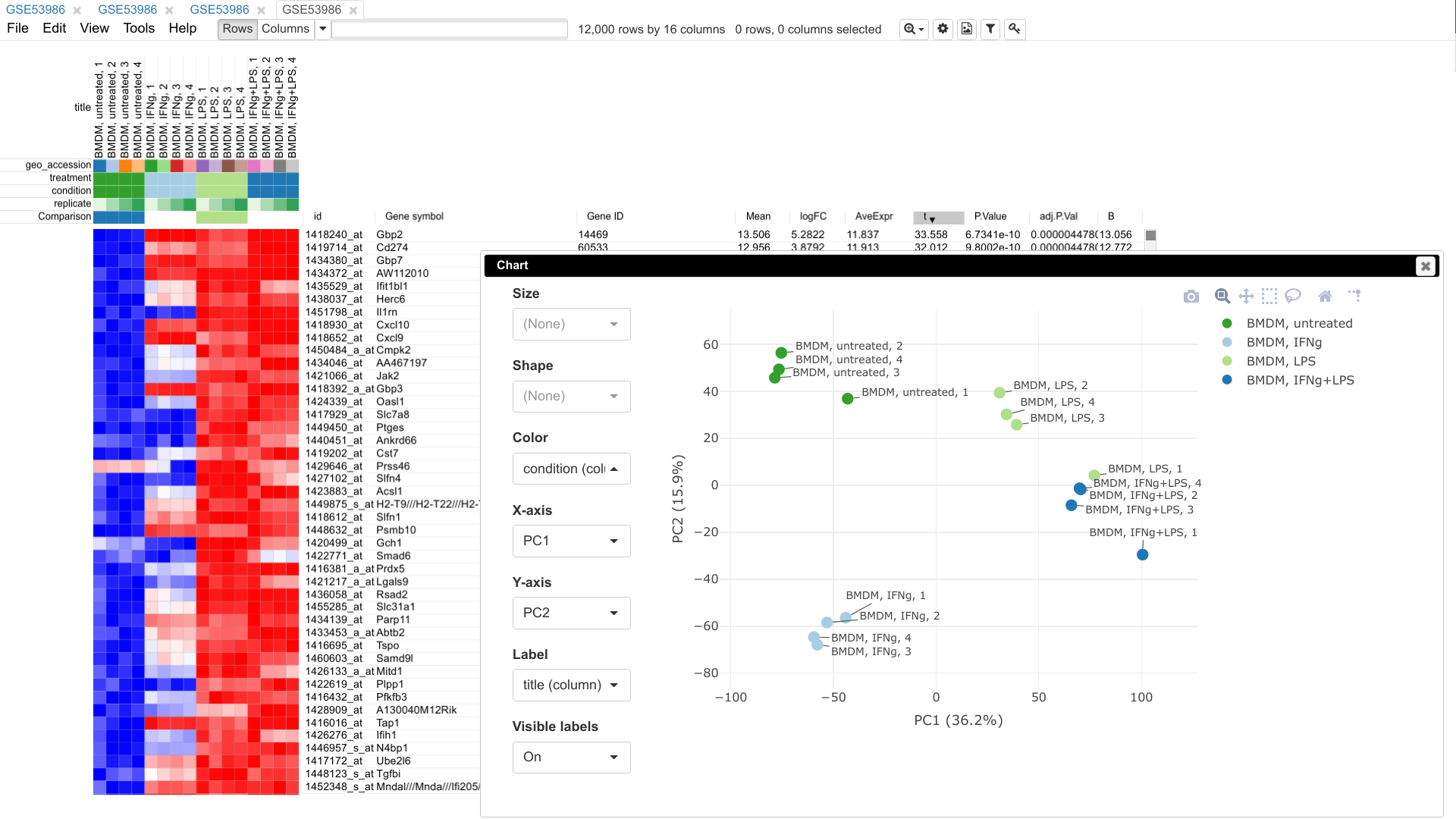
A web tool for visual and interactive gene expression analysis. Integrates an interactive heatmap interface with Bioconductor analysis tools and provides access to >96,000 public datasets.
alserglab/phantasus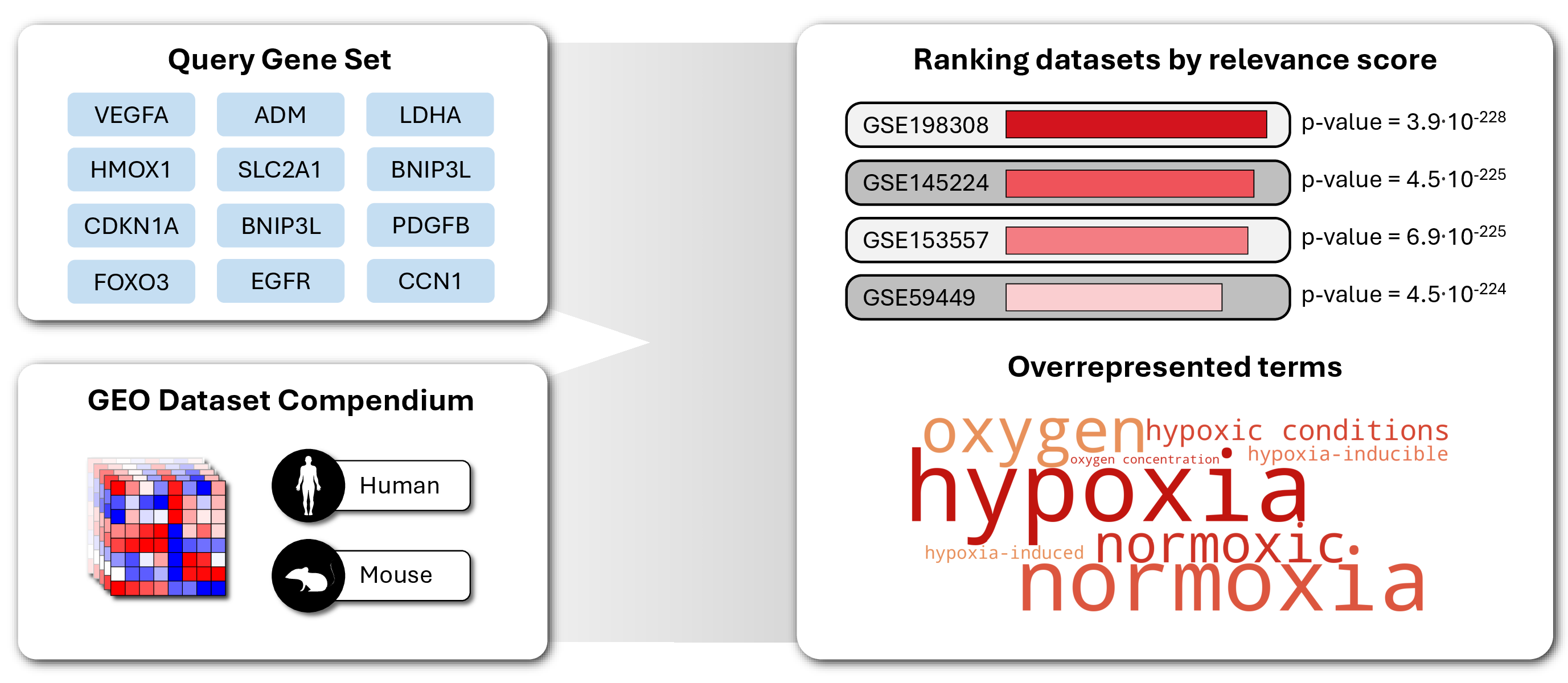
A gene signature-based search engine for public gene expression datasets. Ranks GEO datasets by the level of coregulation of a given gene signature, indexing over 42,000 mouse and 44,000 human datasets.
alserglab/coreshMore
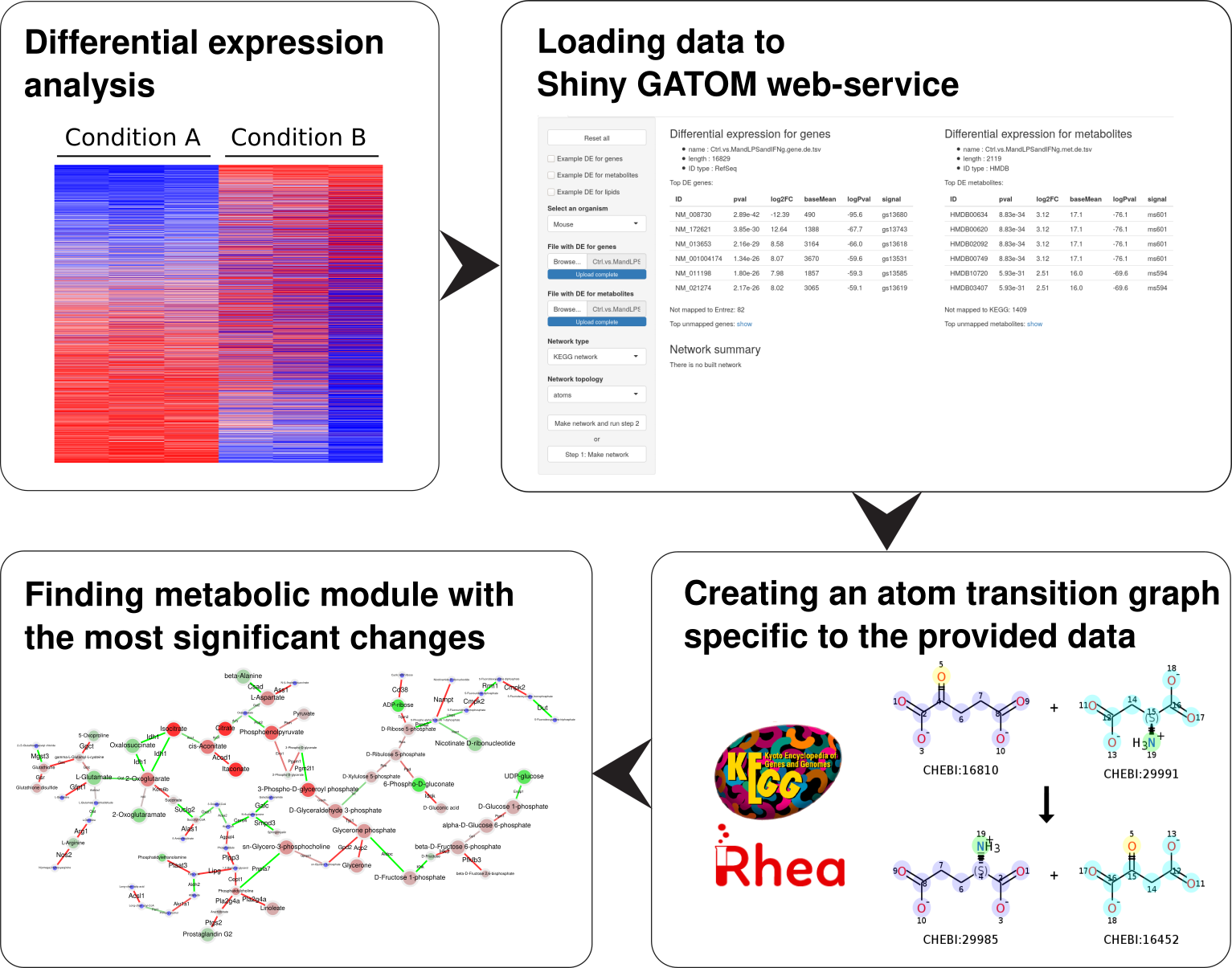
A web service for omics-based identification of regulated metabolic modules in atom transition networks. Supports transcriptional and metabolomic data input, including lipidomics analysis.
ctlab/shinyGatom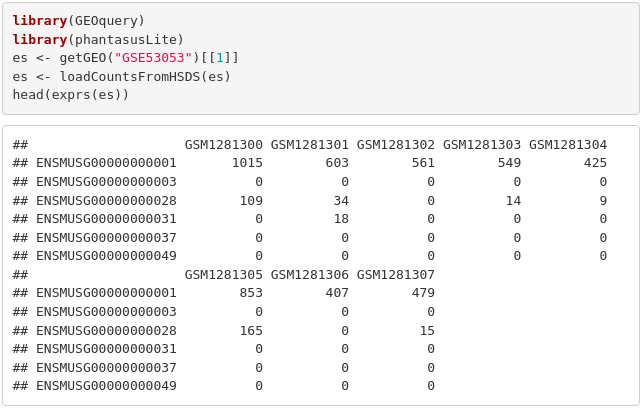
A lightweight R package providing helper functions from Phantasus. Offers an interface to precomputed RNA-seq gene counts from ARCHS4 and DEE2 projects, loading GEO data via GSE identifiers.
alserglab/phantasusLite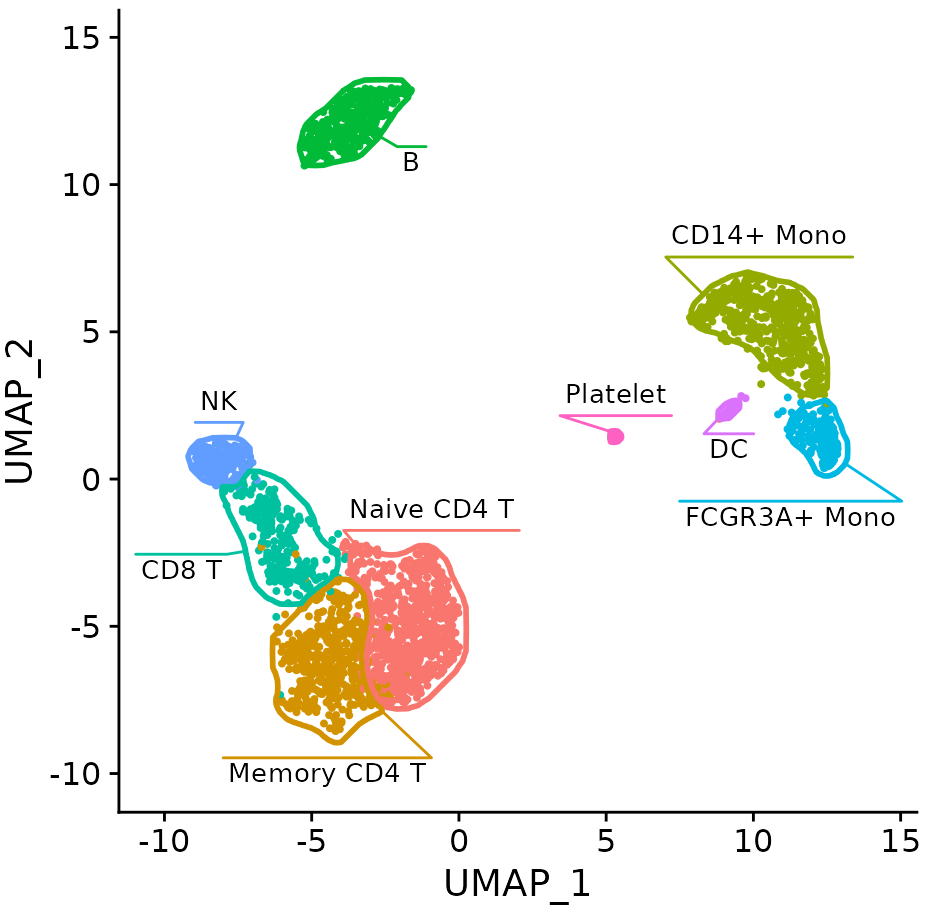
An R package for automatically generating 2D masks for scRNA-seq clusters on t-SNE or UMAP plots. Integrates with Seurat workflows.
alserglab/mascarade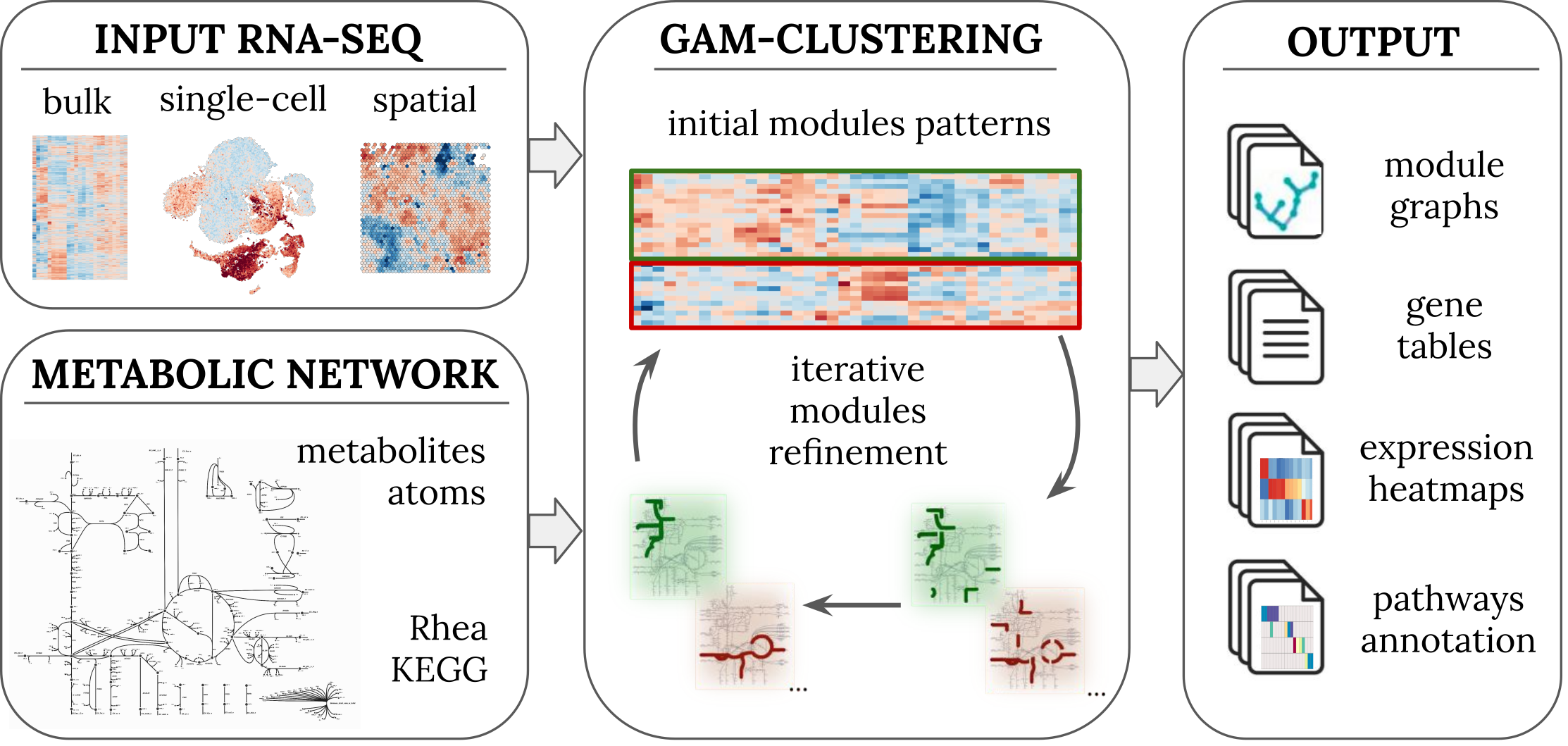
GAM-clustering identifies transcriptionally regulated metabolic modules from RNA-seq data. Works with bulk, single-cell, and spatial RNA-seq.
alserglab/GAMclust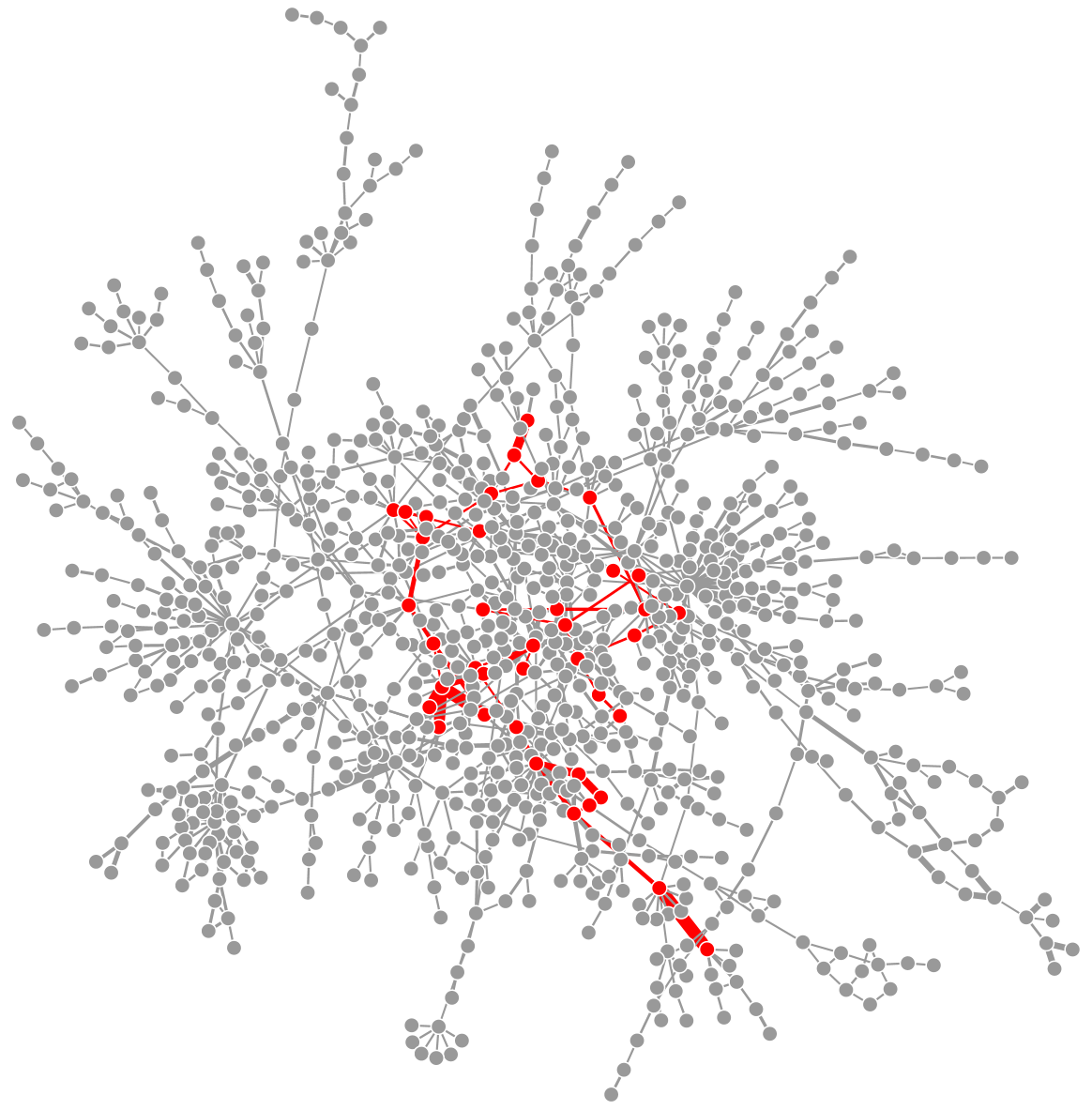
An R package for solving maximum-weight connected subgraph problems and variants. Includes relax-and-cut, simulated annealing, and exact Virgo solvers.
ctlab/mwcsr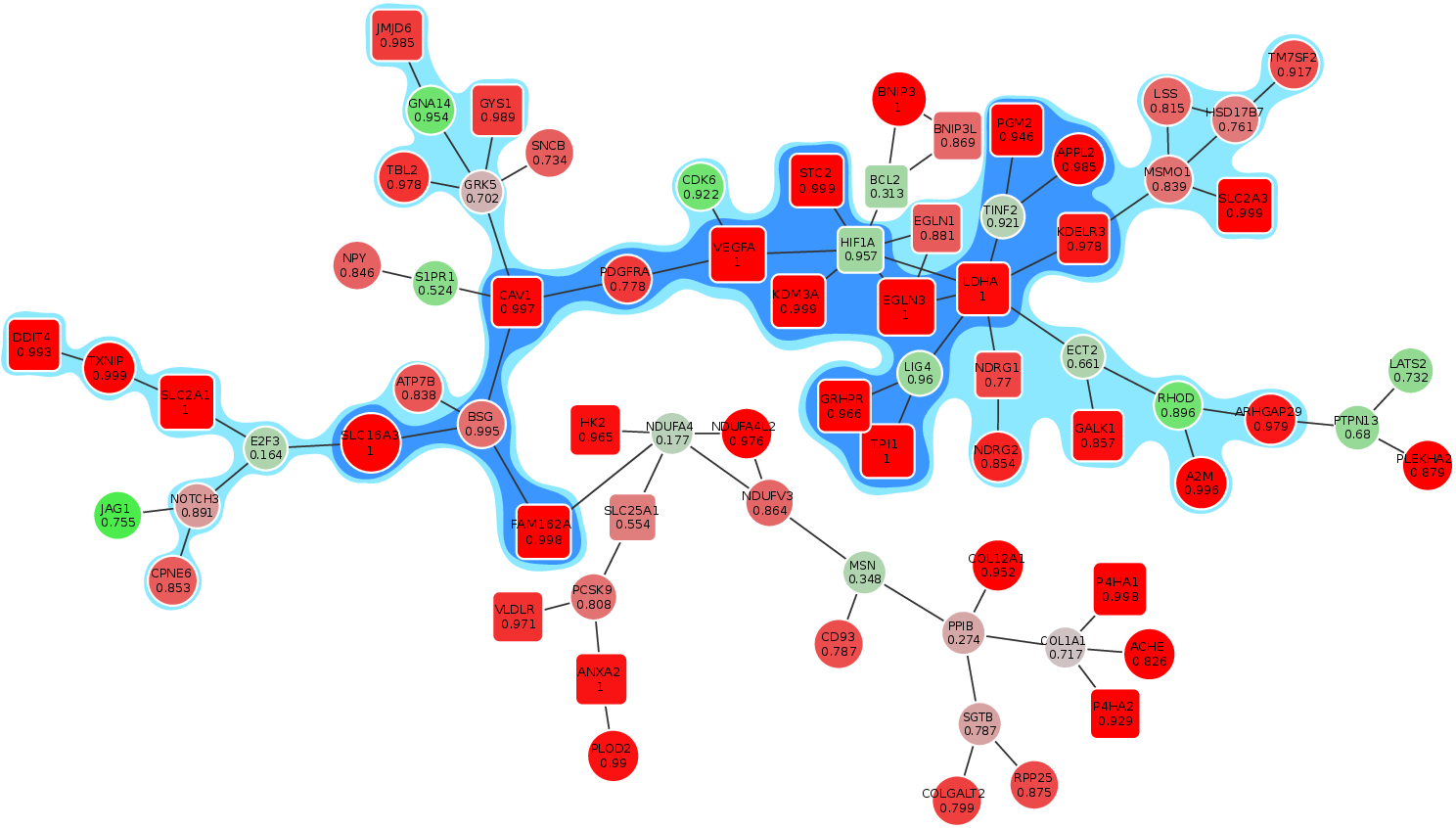
An R package for estimating vertex probabilities in active modules using MCMC methods. Ranks vertices by importance for active network module identification.
ctlab/mcmcRanking