Sergushichev Lab
We develop computational methods and software tools for analysis of omics data, with a focus on transcriptomics and immunology. Our work spans pathway enrichment analysis, network-based integration of multi-omics data, and mining of public gene expression datasets. We aim to make these methods accessible to the broad research community through user-friendly tools.
Research Directions
Pathway Analysis
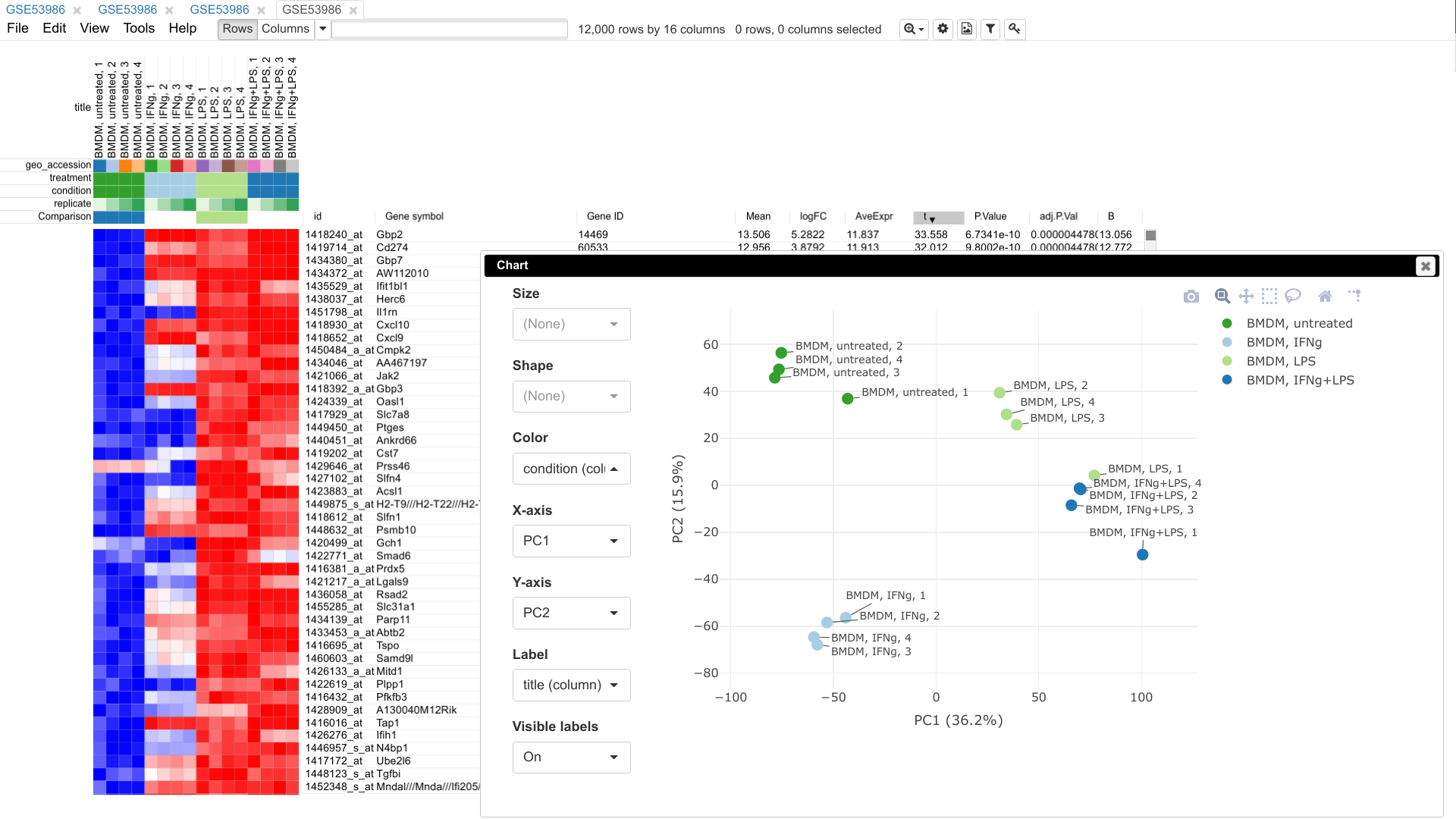
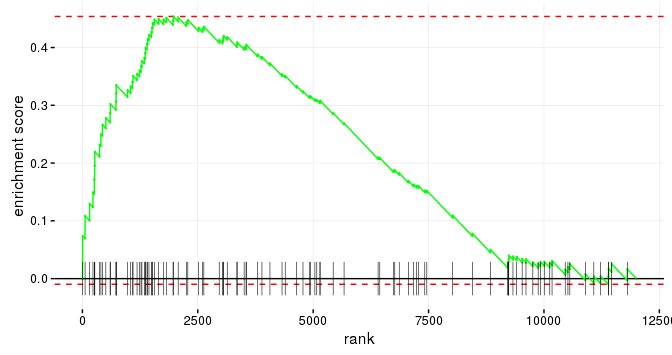
Developing faster and more accurate approaches to pathway enrichment analysis, including methods for modern multi-conditional datasets such as single-cell and spatial RNA-seq. This direction produced FGSEA, one of the most widely used gene set enrichment tools in Bioconductor.
Network Analysis
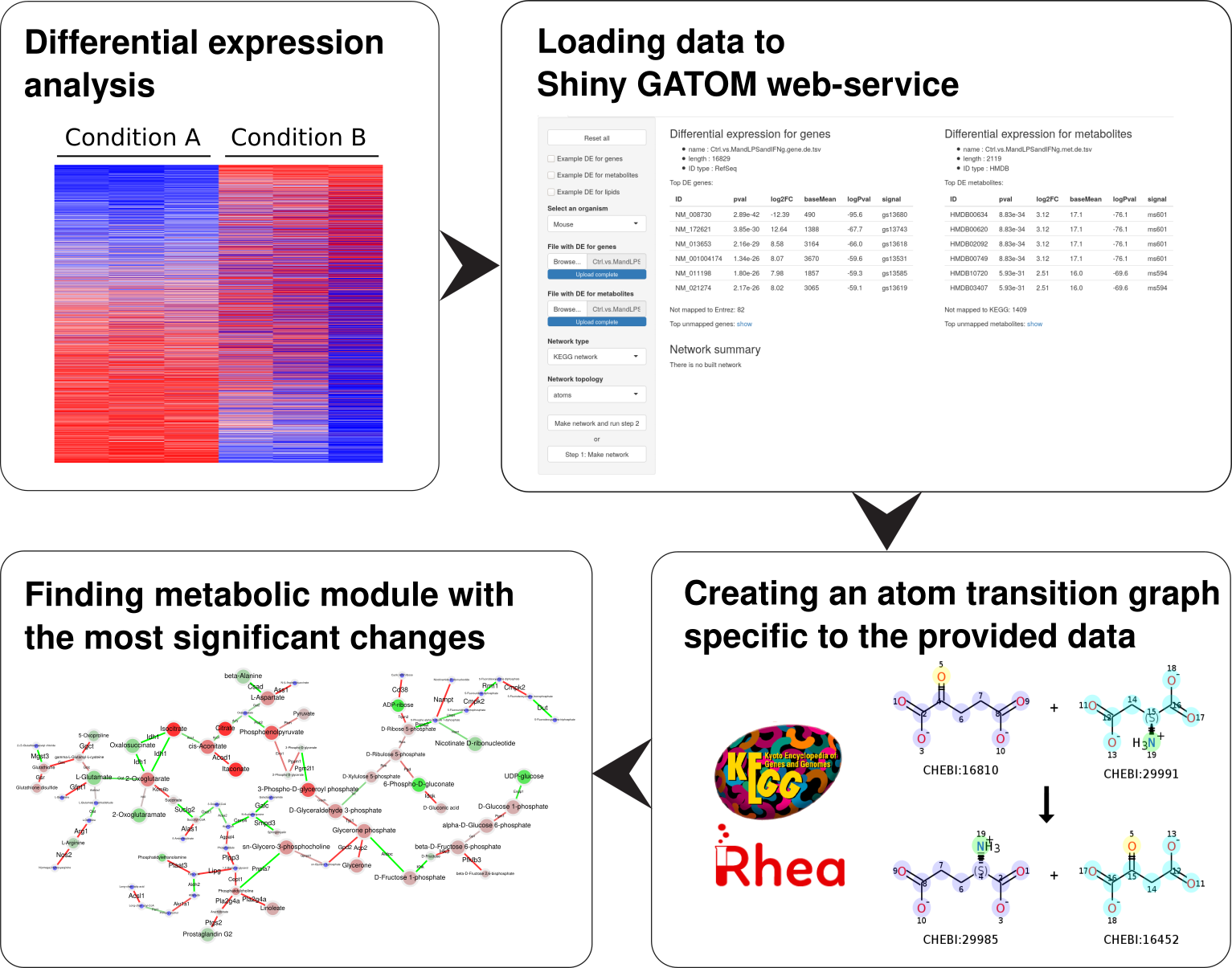
Integrating large-scale omics datasets with biological interaction networks to study cellular metabolic regulation. Our GAM and GATOM approaches use atom transition graphs to identify regulated metabolic modules from transcriptomic and metabolomic data, with applications in immunometabolism.